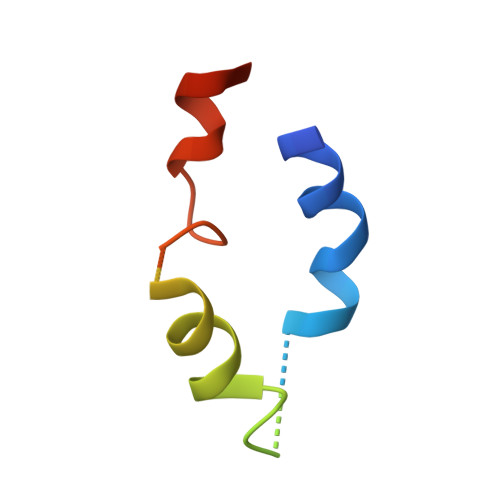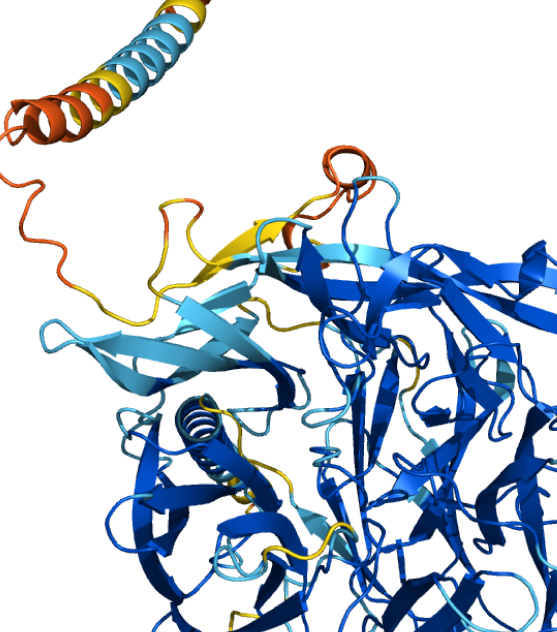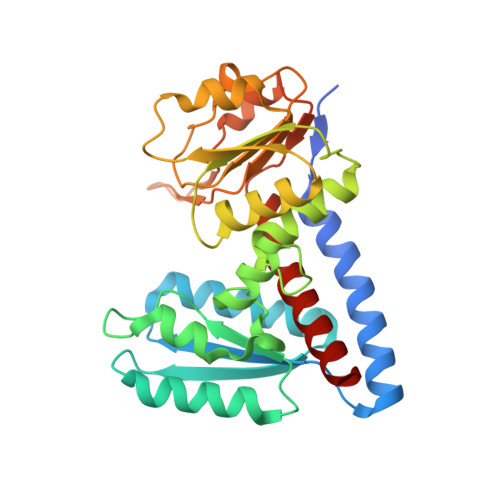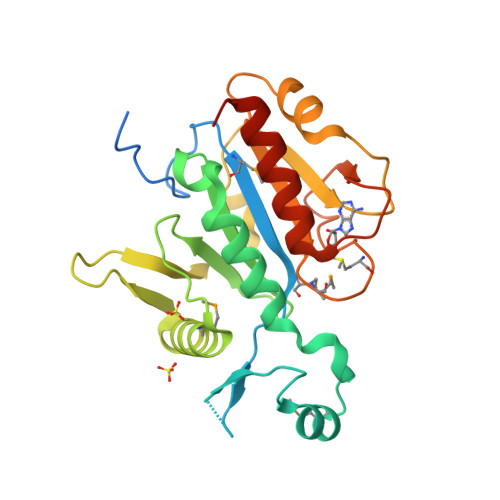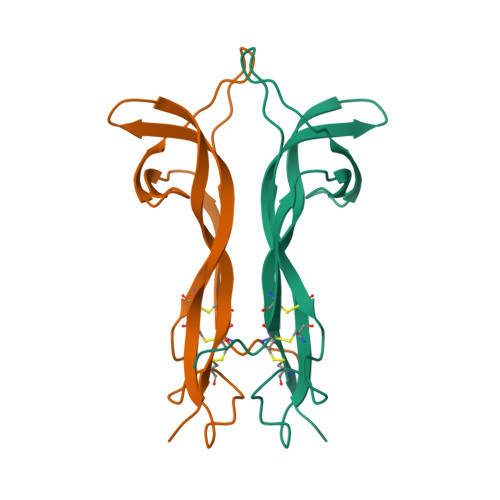
RCSB PDB - 1BET: NEW PROTEIN FOLD REVEALED BY A 2.3 ANGSTROM RESOLUTION CRYSTAL STRUCTURE OF NERVE GROWTH FACTOR
![A 3-D crystal structure of MelB St [pdb access ID, 4M64 Mol-A]. The... | Download Scientific Diagram A 3-D crystal structure of MelB St [pdb access ID, 4M64 Mol-A]. The... | Download Scientific Diagram](https://www.researchgate.net/profile/Markus-Weingarth-2/publication/326807634/figure/fig1/AS:655522487349250@1533300148717/A-3-D-crystal-structure-of-MelB-St-pdb-access-ID-4M64-Mol-A-The-overall-fold-of-MelB.png)
A 3-D crystal structure of MelB St [pdb access ID, 4M64 Mol-A]. The... | Download Scientific Diagram

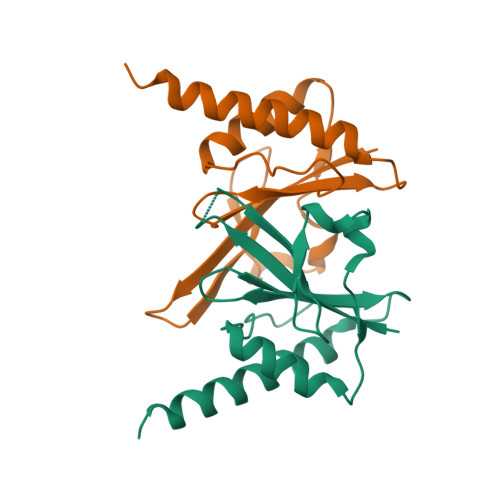
RCSB PDB - 2FDO: Crystal Structure of the Conserved Protein of Unknown Function AF2331 from Archaeoglobus fulgidus DSM 4304 Reveals a New Type of Alpha/Beta Fold

A systematic structural comparison of all solved small proteins deposited in PDB. The effect of disulfide bonds in protein fold - ScienceDirect

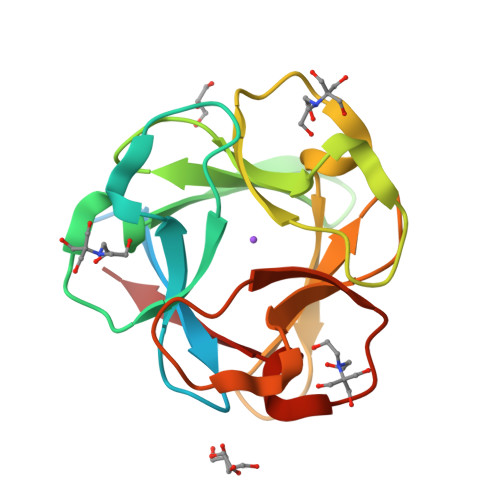
RCSB PDB - 1O70: Novel Fold Revealed by the Structure of a FAS1 Domain Pair from the Insect Cell Adhesion Molecule Fasciclin I

Accuracy of AlphaFold on recent PDB structures The analysed structures... | Download Scientific Diagram

Comparison of the histone fold (PDB: 1HTA)38to eubacteria HU protein... | Download Scientific Diagram
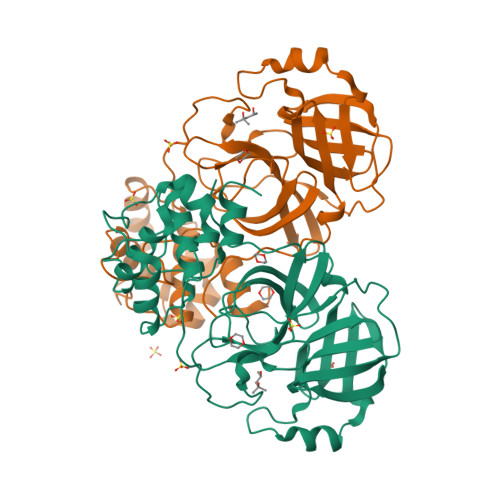
RCSB PDB - 1LVO: Structure of coronavirus main proteinase reveals combination of a chymotrypsin fold with an extra alpha-helical domain
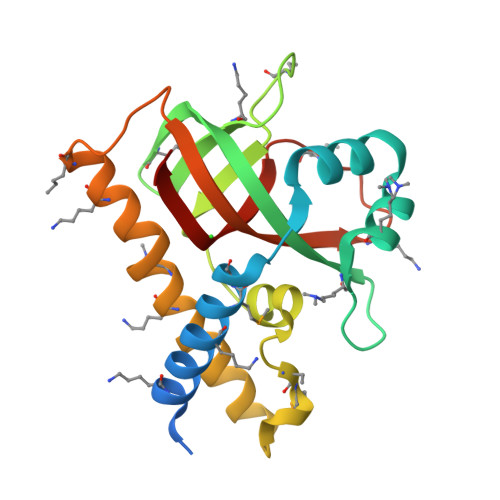
RCSB PDB - 2F4I: Crystal structure of an ob-fold protein (tm0957) from thermotoga maritima msb8 at 1.90 A resolution